Performing
VAI analysis in batch mode
VAI
analysis can also be performed in batch module, like standard perfusion
analysis.
Start the Batch module from the <Modules>
main menu list.
Go to the <Perfusion
analysis> tab and then the <VAI> sub-tab:
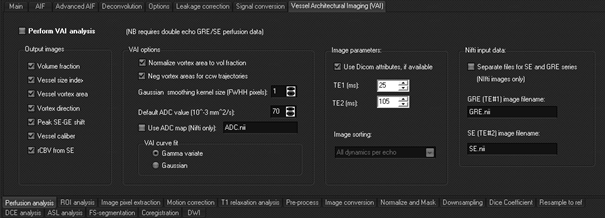
Like
the analysis of standard perfusion data, data can be either in Dicom or Nifti
format. For Dicom images, the VAI series must be present as a single 5D
series containing both the GRE and the SE series in one directory. It is
assumed that the GRE part of the data has lower instance numbers than the SE
series so that the GRE data appears before the SE data upon sorting. For
Nifti input, the data can either be loaded as a single Nifti file containing
both GRE and SE seires, or as two separate files; one for the GRE and one for
the SE series. In the latter case, both files must be present in the same
directory and the same directory as specified in the <perfusion subdir> specified
under the <Main> tab.
An ADC map can also be specified (needed for vessel size index calculation).
The ADC map must be in Nifti format and stored in the same folder as the
perfusion data. If no ADC map is specified, a default constant ADC value (as
specified) will be used.

|